Research
Research
Fibroblast growth factors(FGFs)
Structure and functional investigation of FGF family members with therapeutic potentials
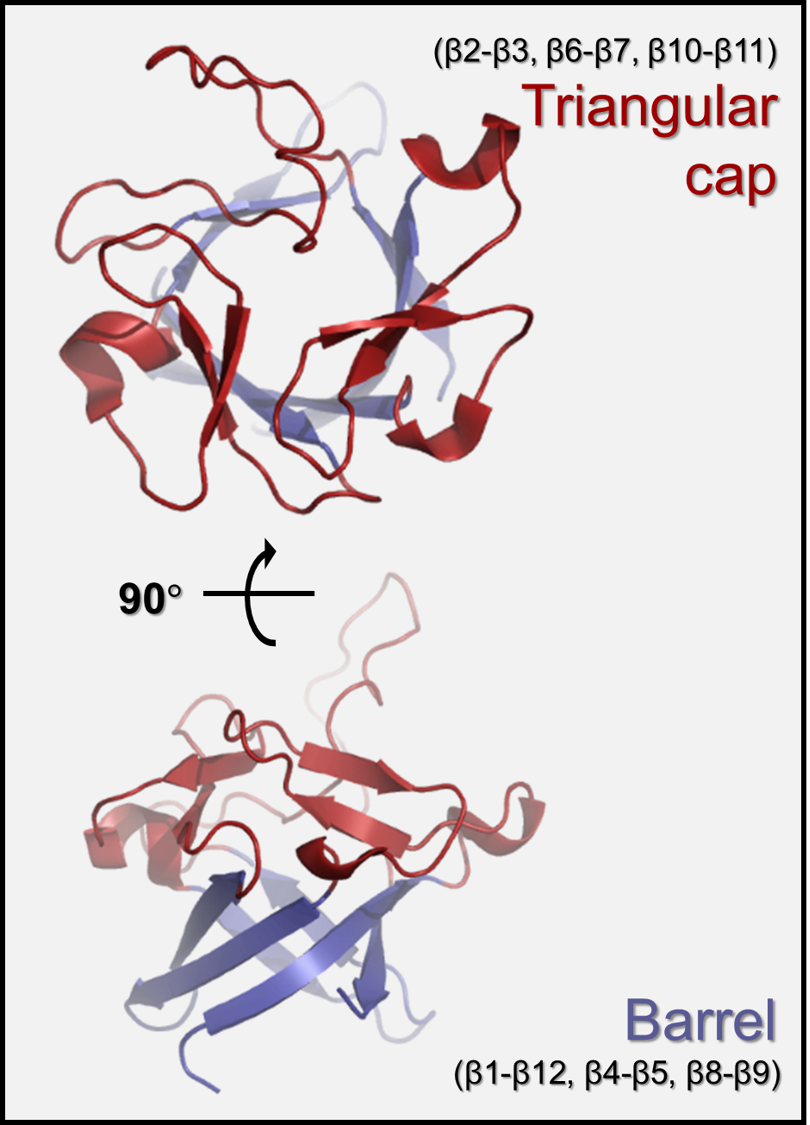
Structure of FGF family
Fibroblast growth factors (FGFs) regulate various developmental and metabolic processes, including cell proliferation, differentiation, angiogenesis, wound healing, nerve regeneration, chronic inflammation, and cancer growth. The FGF family consists of 22 members that share the β-trefoil fold despite their relatively low sequence identities (13–71%). The β-trefoil fold contains 12 β-strands that form 6 two-stranded β-hairpins (i.e., β1-β12, β2-β3, β4-β5, β6-β7, β8-β9, and β10-β11). Hairpins are arranged in a pseudo three-fold symmetry: the first, third, and fifth β-hairpins form a barrel that is covered by a triangular cap consisting of the second, fourth, and sixth β-hairpins.
FGFs are divided into FGF1, FGF4, FGF7, FGF8, FGF9, FGF11, and FGF19 subfamilies depending on their sequence similarities. Except for the intracellular FGF11 subfamily, FGFs form complexes with FGF receptors (FGFRs) on the cell membrane, which are generally composed of three immunoglobulin (Ig)-like extracellular domains, a transmembrane domain, and a cytoplasmic tyrosine kinase domain. FGFs bind to the interface between the second and third Ig-like ectodomains to induce receptor dimerization, leading to the phosphorylation of tyrosine residues in the cytoplasmic domain to stimulate signaling pathways.
The canonical FGFs (FGF1, FGF4, FGF7, FGF8, and FGF9 subfamilies) have a positively charged sector clustered by lysine and arginine residues and thus display avidity for negatively charged heparin/heparan sulfate proteoglycans (HSPGs) present on the cell surface or in the extracellular matrix. Therefore, canonical FGFs form ternary complexes with HSPG and FGFR, and act in the vicinity of cells as paracrine and/or autocrine factors. Conversely, FGF19 subfamily members (FGF19, FGF21, and FGF23) have no apparent HSPG-binding site on their surfaces and thus perform their physiological roles in an endocrine manner. They are released from the extracellular matrix and reach remote target organs through the bloodstream.
[Related publications]

- Young Jun An¶, Ye-Eun Jung¶, Kyeong Won Lee, Prashant Kaushal, In Young Ko, Seung Min Shin, Sangho Ji, Wookyung Yu, Cheolju Lee, Won-Kyu Lee, Kiweon Cha, Jung-Hyun Lee*, Sun-Shin Cha*, Hyung-Soon Yim*
"Structural and biochemical investigation into stable FGF2 mutants with novel mutation sites and hydrophobic replacements for surface-exposed cysteines" - PLoS ONE, Jul 08; accepted (2024)
Ye-Eun Jung¶, Kyeong Won Lee¶, Jae Hyun Cho¶, Da-Woon Bae, Bo-Gyeong Jeong,Yeon-Ju Jung, Soo-Bong Park, Young Jun An, Kyungchan Kim, Ga Seul Lee, Lin-Woo Kang, Jeong Hee Moon, Jung-Hyun Lee, Eun-Kyoung Kim*, Hyung-Soon Yim*, and Sun-Shin Cha*
"Heating-mediated purification of active FGF21 and structure-based design of its variant with enhanced potency"
Scientific Reports, Jan 18;13(1):1005 (2023)
Kyeong Won Lee¶, Young Jun An¶, Janet Lee¶, Ye-Eun Jung, In Young Ko, Jonghwa Jin, Ji Hoon Park, Won Kyu Lee, Kiweon Cha, Sun-Shin Cha, Jung-Hyun Lee*, Hyung-Soon Yim*
"Expression and purification of intracrine human FGF 11 and study on its FGFR-dependent biological activity."
Journal of Microbiology, Nov 01;60(11):1086-1094 (2022)
Young Jun An, Kyeong Won Lee, Ye-Eun Jung, Ye Eun Jeong, Su-Jin Kim, Jurang Woo, Jonghwa Jin, Won-Kyu Lee, Kiweon Cha, Sun-Shin Cha, Jung-Hyun Lee*, and Hyung-Soon Yim*
"Improvement of FGF7 thermal stability by introduction of mutations in close vicinity to disulfide bond and surface salt bridge"
International Journal of Peptide Research and Therapeutics, Mar 29;28:85 (2022)
Julee Kim, Sangki Baek, Yean Ju Hong, Michelle Novais de Paula, Musharrat Jahan Prima, Yeon-Mok Oh, Sun-Shin Cha, Jeong Tae Do, Yeon Jin Jang, Han Choe*
"DJ-1 can replace FGF-2 in defined media and feeder-free condition for long-term culture of human pluripotent stem cells"
International Journal of Molecular Sciences, May 31; 22(11):5954 (2021)
Kiwoong Nam, Kyeong Won Lee, Oksung Chung, Hyung-Soon Yim, Sun-Shin Cha, Sae-Won Lee, JeHoon Jun, Yun Sung Cho, Jong Bhak, Joao Pedro de Magalhaes, Jung-Hyun Lee*, Jae-Yeon Jeong*
"Analysis of the FGF gene family provides insights into aquatic adaptation in cetaceans"
Scientific Reports, 7:40233 (2017)

